[Note to readers: This post is far longer than the any I have previously published on this blog. It had to be: it is about a complex topic, and it attempts to correct a significant historical oversight. I encourage you to read to the end, skipping the more technical parts if you wish. — G]
We’ve all heard about the race to sequence the human genome that started in 1990 and culminated with the announcement in 2003 that the sequence was completed. This achievement was accomplished by two organizations (one private and one public), and enabled by the work of thousands of scientists using hundreds of sequencing machines.
What you may not have heard is that two of our Kendal residents, Jim Prober and Charles Robertson, played a leading role in developing those sequencing machines.
Sequencing the human genome is one of the great scientific milestones of our age. Among other things, sequencing the human genome has made it possible to identify the causes of many diseases, to reveal many of the mechanisms by which our bodies operate, to show the details of how heredity works, to prove the identity of criminals, and much more.
As one example of the medical importance of sequencing, consider cystic fibrosis (CF). Most CF sufferers have a mutation in the gene for a critical protein (called CFTR) that controls the passage of ions through the cell wall. The mutation results in a faulty protein that does not have the right shape. In the lungs, this misshapen protein can cause mucus to accumulate, with serious consequences. Based on this knowledge, medications for CF were developed.
So how does that example relate to DNA sequencing? All DNA is composed of just four subunits (“bases” or “nucleotides”): adenine (A), thymine (T), guanine (G), and cytosine (C). The “human genome” is a listing, in sequence, of the billions of bases in the human chromosomes, which are the hereditary material contained in every human cell. The mutation responsible for most CF cases in people of European heritage can be traced to just three missing bases at position 508 of the DNA for the CFTR protein, as shown in the diagram below.
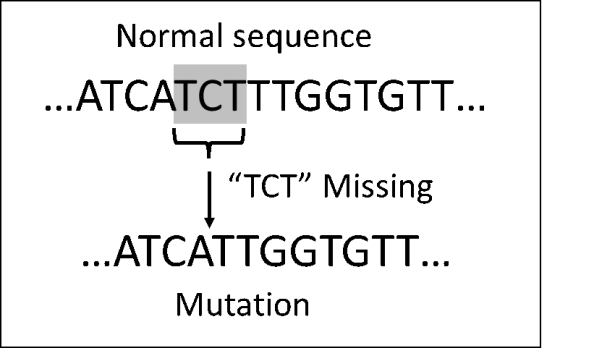
This blog post is about the sequencing machines that made this type of knowledge possible. Standard accounts in books and on the web claim that most of those machines came from a company called Applied Biosystems (ABI), which developed them based on work done by researchers at Cal Tech.
That’s the usual story, but it is not entirely accurate.
It is true that ABI machines dominated the sequencing of the human genome. But arguably, the technology that first demonstrated how DNA sequencing could be efficiently automated was not from ABI or from Cal Tech. It was invented by a small group, in a little-known, under-the-radar project at DuPont. Two Kendal residents, Charles Robertson and Jim Prober, played critical roles in that project.
The key question that the Human Genome Project set out to answer was: What is the sequence of the A, C, G, and T bases in the chromosomes of a human being? Each human cell contains 3 billion pairs of bases, so the task of sequencing them all was never going to be easy.
By the early 1980s, when Jim and Charles first got interested, very little of the human genome had been sequenced. The only genomes completely sequenced were those of some viruses, bacteria, and a few other very simple organisms.
[Note: Feel free to skip down to the next section, “How DuPont got involved,” if you are not interested in the details of how sequencing was done prior to the DuPont project. ]
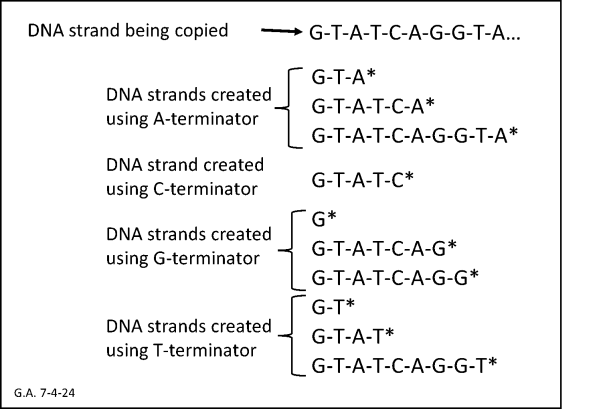
Sanger sequencing. In the early 1980s, the quickest way to sequence a gene was called “Sanger sequencing.” Developed by Fred Sanger at Cambridge (UK), this approach used a well-established technique for making copies of a specific stretch of DNA in a test tube. Sanger’s breakthrough was developing a set of chemical agents that could stop the growth of DNA copies in their tracks at a specific type of base (A, C, G or T). Let’s call these agents “A-terminator,” “C-terminator,” “G-terminator,” and “T-terminator” (the official name is “ddNTPs”). Sanger would take one test tube in which a particular stretch of DNA was being copied and add a tiny amount of A-terminator. Soon, pieces of DNA of various lengths would start accumulating as their growth was blocked by an A-terminator incorporated instead of a normal A-type base at the end of the growing chain.
Suppose the stretch of DNA being copied had the sequence of bases GTATCAGGTA. By looking at the sequence, you can see that the A’s are in positions 3, 6, and 10. The A-terminator would stop the DNA copying at those positions, and the A-terminator test tube would end up with many copies of DNA that were 3, 6, and 10 bases long, but none that were 1, 2, 4, 5, 7, 8, or 9 bases long. Sanger could sort the pieces by length using a technique called “gel electrophoresis”, in which shorter pieces moved through a gel more quickly than longer ones. This process was repeated for all four terminators, in four parallel “lanes” in the two-dimensional gel. The result was a slab of gel with the strands of DNA spread out along each lane according to their length, corresponding to one of the four bases.
By adding a radioactive tracer to the strands of DNA and exposing the gel to x-ray film, Sanger got a black band on the film at each location where a cluster of strands of DNA ended up. The distance they had moved through the gel (their “mobility”) told him the length of the strands in each cluster. By comparing the distances travelled through the gel by all four types, he could determine the sequence of bases in the specific stretch of DNA he was working on.
Sanger’s revolutionary technique earned him a share of the 1980 Nobel Prize. It worked well, and it was faster than previous approaches. But it required using radioactive chemicals, and it took several days and considerable effort by a human reading the resulting x-ray films to obtain the sequence of only a few hundred base pairs. That process was subject to errors and could not be scaled up to the level needed for the three billion base pairs of the human genome. It also generated a large amount of radioactive waste, which was a problem for disposal.
How DuPont got involved. Jim Prober’s office mate at the Dupont Engineering Physics Lab, Rudy Dam, had a background in physics and molecular biology, and wanted to explore whether the Sanger process could be automated. In early 1983, Rudy signed up for a biotech conference in San Diego and Jim decided to tag along. They checked out the state-of-the-art equipment. After learning the details of the Sanger process, they concurred that it was insufficient and that a totally automated approach would be needed to sequence a complex genome. They figured they could do better.
On their return to DuPont, Jim and Rudy talked the situation over with Charles Robertson, a gifted and creative research engineer that worked in the same lab. Initially, they did not seek input from management. They thought it was prudent to stay “under the radar,” at least for the initial stages of the project.
With Charles on board, they had the core of a powerful team. In addition to the background Jim and Rudy had in biology, physics, and integrating complex instrument systems, they knew about the political workings of DuPont and how to navigate the corporate structure to keep the project out of trouble. Charles brought outstanding expertise in optics, mechanics, and electronics.
The team needed to develop a system that would be automated, more accurate, and much faster than the current state of the art. They agreed that it wasn’t practical to automate the Sanger process. That would have required automating the detection of tiny amounts of radioactivity, skipping the x-ray film step. The technology did not exist to do that, so the team considered alternative approaches.
After some brainstorming, they decided to investigate a detection method that used fluorescence instead of radioactivity.
If they could figure out how to attach a different fluorescent “tag” (an arrangement of atoms that emitted light when illuminated by a laser) to each of the DNA fragments ending in A, C, G, or T, then they could avoid the four separate gel “lanes”, one for each type, required for the Sanger process. A single lane would do, with the identity of the fragments being known, not because it was in a certain lane in the gel, but because of the color of the fluorescence.
[Note: The following section can be skipped if you are not interested in how the Sanger process was adapted to fluorescent dyes. The story resumes at “Obstacles and slow progress”.]
Using fluorescent dyes: a promising idea, but could they make it work? The key to success was a set of four “fluorescent terminators,” meaning ddNTPs that are coupled to fluorescent dyes instead of radioactive tracers. Each terminator would display a distinguishable fluorescence pattern when illuminated by a laser. As in the Sanger method, such terminators would be incorporated as part of the process of terminating the growing DNA chain at a specific base. And, importantly, each terminator had to have nearly the same size, shape, and charge so that the attached DNA strands would move through the gel at a pace that was strictly proportional to their length. The speed had to be independent of whether the strand had an A, C, G, or T terminator. In essence, the strands would assume an unchanging “marching order” based on their length, as they moved through the gel.
Instead of the dark bands on the X-ray film that Sanger obtained, the fluorescence technique would detect moving patches of different-length strands, distinguishable by the colors of their fluorescence. And instead of Sanger’s four separate test-tube mixtures, all the dye attachment chemistry could be done in one tube, which would be a great advantage when scaling up to large sequencing projects. By using a laser to trigger the fluorescence as the stands of DNA moved through the gel, it would be possible to detect the size of the strands, and therefore the DNA sequence, in real time. These features would comprise the basis of a completely automated process, suitable for scale-up and commercialization.
Obstacles and slow progress. In principle, the fluorescence approach certainly looked promising, but there were huge challenges. To turn the concept into reality, the Dupont team had to overcome major technical obstacles. These included developing the right set of four fluorescent dyes; creating the dye-tagged terminators that efficiently and accurately incorporated into the growing DNA strand; and building a sensitive and reliable detection system.
By this time, the team recognized that they needed someone who understood the Sanger process and was an expert in biochemistry and fluorescence. A senior DuPont manager, Vinay Chowdhry (who still lives near Kendal), set up a meeting to interview potential DuPont employees who might fill that role. One candidate stood out: an organic chemist named George Trainor. Though he recognized it would be a major challenge, George responded enthusiastically, and felt confident that the right dyes, and a method of attaching them to the DNA strands, could be developed. Shortly after that meeting, Trainor joined the team.
Jim Prober and Rudy Dam (with Charles Robertson adding his engineering input) went to work on the design of the sequencing process and the dye detection technology. Meanwhile, George Trainor began work on the fluorescent dyes and the challenge of attaching them to the strands of DNA.
Ideally, the team would have preferred four dyes with fluorescence that covered distinct parts of the light spectrum. That would have made detection and identification with simple light filters relatively easy. But they soon realized that was unworkable. Dyes with those properties would be too different in size, shape, and charge to keep the tagged DNA strands in a reliable length-based “marching order” in the gel. The various lengths of DNA would become intermingled, making detection of the sequence impossible. Instead, Trainor set to work on a set of four chemically similar dyes with partially overlapping emission spectra (see diagram below).
On the molecular biology front, the DuPont team struggled to find a way to attach the fluorescent dyes to the strands of DNA. The combination of terminator plus dye had to be compatible with DNA polymerase, the enzyme that attaches bases to a growing DNA strand. Frank Hobbs, a chemist working with George Trainor, developed a “linker arm” concept that would serve as a bridge to attach the fluorescent dye to the end of the DNA strand. They tried one linker arm structure after another, but success was elusive. The structures interfered with the operation of the DNA polymerase (by design, enzymes are very unforgiving), and the dyes failed to incorporate into the DNA strands.
Test after test. This work proceeded slowly, by trial and error, with numerous linkers being tried over a period of many months. During this phase, Trainor and Hobbs, who by now had assembled a small team of co-workers, focused on developing a workable version of the fluorescent “T-terminator,” one of the four that would eventually be required. They assumed that if one terminator worked successfully, they could create the other three, hopefully with less effort. While deeply challenged, Trainor and his team never lost confidence that the problem would eventually be solved.
A testing routine developed. Roughly every few weeks, when a new terminator/linker arm candidate became available, the chemistry team delivered a DNA sample for a run through the test apparatus. A voltage was applied to the sequencing gel, the sample was injected at the top, and the wait began. If the terminator was successful, the sample would move past the laser fluorescence detector in ten minutes or so, and a strip-chart recorder would show a peak.
That’s what was supposed to happen, but time after time, there was no peak. Jim, as the team member living closest to the lab, got the job of staying late to run the tests. Over a period of months, each candidate was tested, each one failed, and Jim headed home with no news to report.
One memorable evening, a candidate linker arm configuration finally worked. When the peak on the strip-chart started to appear, Jim held his breath. Would it turn out to be a small, random squiggle? No, it was a strong, clear peak. When it happened, Jim was so excited he called up people all over DuPont—team members and leaders—to spread the news. That breakthrough finally proved that the fluorescent approach could work.
Success at last. Although it would take about a year before the three remaining dyes and linker arms were successfully synthesized, the team now had a clear path ahead. George, Frank, and their team created a set of four closely-related dyes with overlapping spectra—not as distinct in color as they would have liked—but which could be distinguished from each other by their fluorescence signals at two carefully-chosen ranges of wavelengths. The dyes had very similar chemical structures, so they had compatible characteristics in the electrophoresis process. And, crucially, the linker did not interfere with the attachment of the dye to the DNA strand.
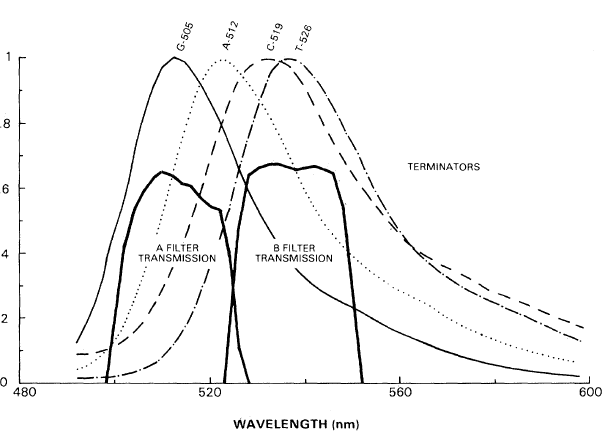
Meanwhile, Charles had developed the detection system, comprised of a laser, filters, and other optical components to stimulate and detect the fluorescence, along with the associated electronics and the strip-chart recorder for output. He chose the filters to isolate the two ranges of wavelengths used to distinguish the fluorescent tags; the ratio of the two signals identified each base as it passed the detector. Charles made use of rectangular filters that could be placed flat against the glass surface holding the gel, along with a photomultiplier tube (to detect the fluorescence) the same shape as the filters. This arrangement maximized the signal generated by the fluorescent tags.
While this work was going on, interest in the human genome was ramping up in the scientific community. The need for much more efficient sequencing was becoming clear. The timing was perfect for the DuPont team.
When the team successfully demonstrated the fluorescence sequencer concept, other groups at DuPont (including some who had previously disparaged the effort) jockeyed for the opportunity to take over the project. But with the support of DuPont upper management, the team was able to continue its development work without interference.
A working model. Once all the key elements of the system were working, prominent people in the DNA sequencing field were invited to the lab to see the equipment. The demonstrations worked, but the parts were spread out on a lab bench about half the size of a ping-pong table. Still, many of the visitors, including Nobel prize-winners Sydney Brenner from Cambridge (who was already acting as a consultant on the project) and Harvard’s Walter Gilbert saw the potential and were very excited.
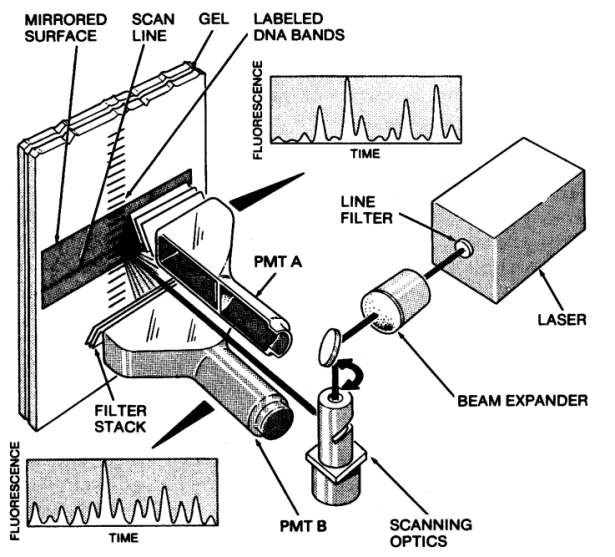
The push for a commercial product. Word of the sequencer spread within Dupont, and there was again an effort by another DuPont group to take over the project. An emissary arrived, with the message that the team was to hand the sequencer project off to his group. “You guys have done great work, but your expertise is too important to keep working on this,” he said, “We’ll take it from here.”
But the team knew that approach would never work, given the stage that their project was in. They still had much work to do before the sequencer could be entrusted to a commercial development group. Jim stated what they all recognized: it was time to start work on a prototype commercial product of their own, and that’s what they did. They knew that some key DuPont executives were quietly supporting their work and would provide cover going forward.
Charles immediately set about creating a design that could lead to commercialization. That meant corralling all the components scattered on the lab bench (laser, detection equipment, gel column, and associated gear) into a compact desktop instrument that could be operated without too much training and be repaired in the field. In a matter of days, Charles drew up the plans for assembling and packaging such a device. Then he contacted a company he knew of in Philadelphia that could design and build a professional-looking mockup of the instrument housing. They quoted him a figure of $42,000 for the work.
$42,000 was a lot of money in the mid-1980s. Jim didn’t have the authority to make a $42,000 purchase. He took a deep breath and signed off on the $42,000 anyway. The authorization went through, which was clear evidence of corporate support.
By late 1986, the mockup was almost ready. And the timing was perfect. At the end of the year, the team was invited to show their work to DuPont’s executives. They took the sequencer mockup (still smelling of fresh paint) to a special exhibition room covered with a drape and surrounded by a series of stations explaining the underlying technology. At the right moment in the presentation, the drape was removed, revealing the new instrument. The presentation was enthusiastically received, and the team knew they had the green light to proceed.
To fully automate the sequencing process, a small computer would be needed to function as an instrument controller. At the time, the obvious choice would have been the IBM PC. But the new Macintosh desktop model (Macintosh II) had just come out. It was powerful and looked great. The user interface was way more elegant and user-friendly than the IBM PC. The introduction of the Macintosh II made the front page of USA Today. The team wanted to use it in order to take advantage of the excellent look and feel the Mac II provided. The DuPont IT department had an Apple connection, and Apple immediately sent an executive team to meet with the DuPont team. After brief discussions, Apple agreed to support the project. This was the first opportunity for Apple to expand beyond personal and business computing into scientific instrument control. DuPont had only limited abilities to develop software for the Macintosh II, so Apple arranged to bring in a manager and three graduate students from Drexel University to work on the software. Jim suddenly found himself in charge of software development.
Three commercial prototypes were built, and the instrument was christened the Genesis 2000. At that point, the development team felt they had taken the project as far as they could. A new team was needed for commercialization. Management assigned a seasoned manager, Karl Jahn, to take the instrument to market. Karl, who had a successful track record of commercializing products in the biotech industry, immediately became committed to the success of the project. Rather than a “we’ll take it from here” approach, he consulted closely with Jim, Rudy, and Charles to learn the details of the system and how it represented a revolutionary approach to sequencing. He used this input to organize the commercialization effort. This included carefully selecting new team members that could scale up production, testing and quality control, and building a marketing and sales campaign. Jim, Rudy, Charles, and the other players who built the first prototypes remained key members of the team. Working in a new facility dubbed the “skunk works,” the commercialization and marketing team built and sold about 125 units into the field.

The aftermath. DuPont did not have field technicians everywhere that could manage the Genesis 2000 system. This limited where and to whom the sequencers could be sold. The DuPont team could not begin to keep up with the demand. They settled on a policy of only selling them to “opinion leaders”—the top people in the DNA sequencing field, including leading participants in the Human Genome Project. Eventually, DuPont sold the technology rights to Applied Biosystems (ABI). ABI went on to sell thousands of sequencers, based on the DuPont fluorescent terminator technology and subsequent improvements.
Although the DuPont fluorescent dyes and detector approach were replaced in ABI machines by 1994, I am told that the linker chemistry is still used to this day.
The DuPont team received widespread recognition for their work when they published a paper about it in Science, the premier US scientific journal. It was the cover story of the October 16, 1987 issue. Jim and Charles were among the nine authors listed (so were Rudy, George, and Frank). Jim was the lead author and Charles provided important artwork, including a detailed diagram showing the workings of the machine.
DuPont applied for and received several patents on the sequencer technology. The main patent covering the workings of the machine itself (patent number 5306618) was granted in 1992. Jim and Charles were listed among the five “inventors” (the others being Rudy, George, and Frank).
I think it is safe to say that the DuPont project provided the breakthrough that first made it possible to imagine that sequencing the entire human genome might be feasible. The Human Genome Project officially began in 1990. By the time the sequencing of the genome was completed, in 2003, most of the sequencing had been done on the Genesis 2000 and its successor machines from ABI. It’s fair to say that Charles, Jim, and their team at DuPont played a crucial role in one of the great scientific achievements of our lifetime. I feel privileged to know them.

Absolutely fascinating, although I understood very little of the process. George is amazing at being able to describe it for us. Kudos and thanks to Jim and Charles. Wish I had even a tiny understanding of this complex field!
LikeLike